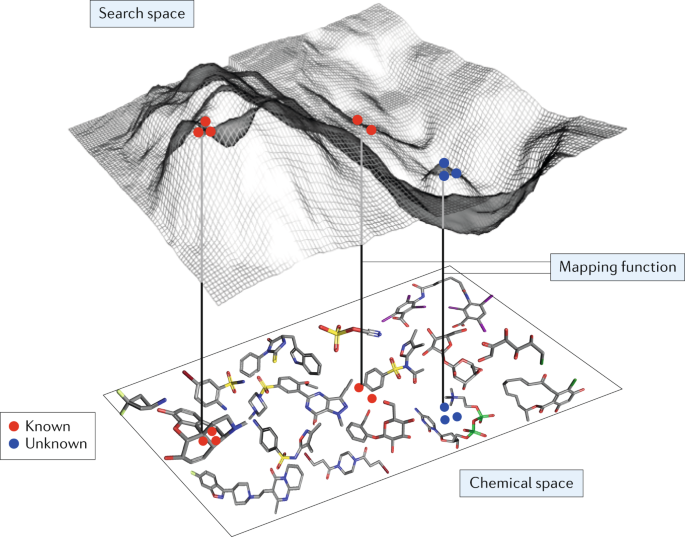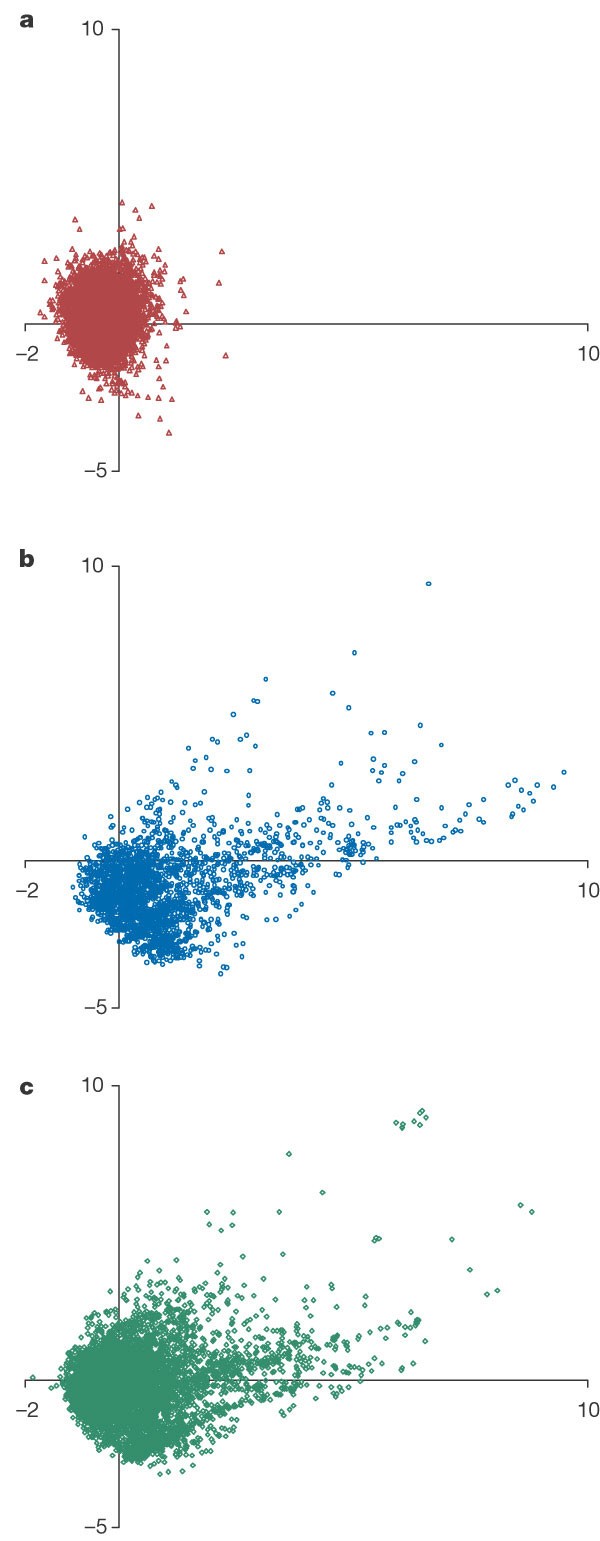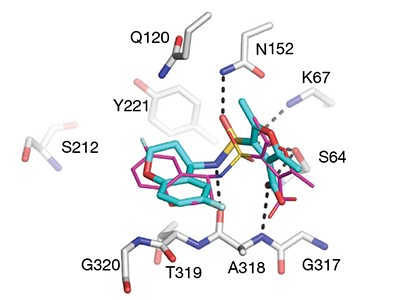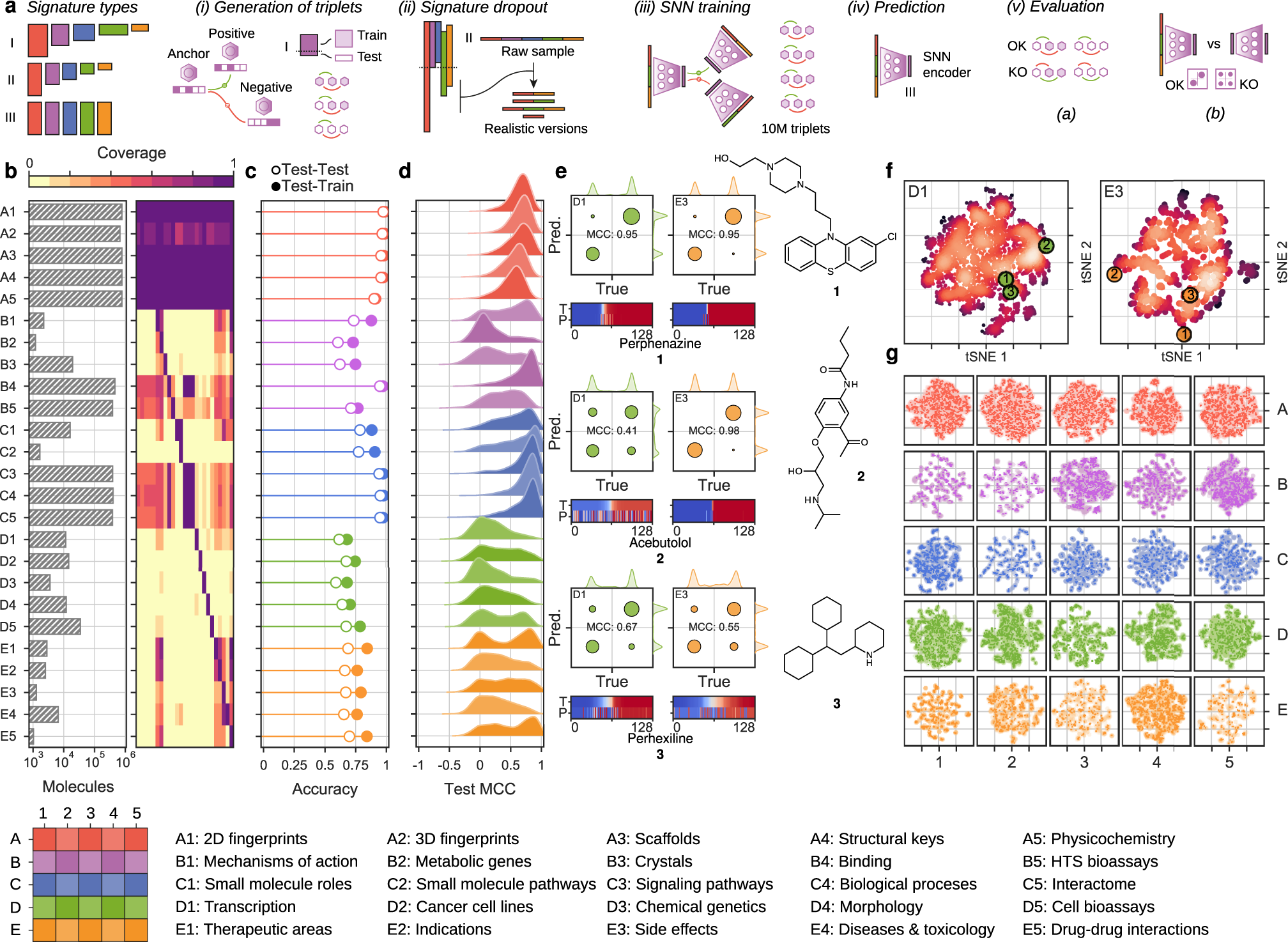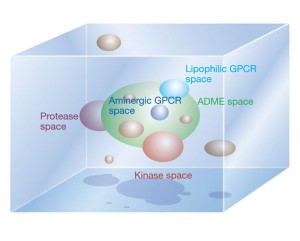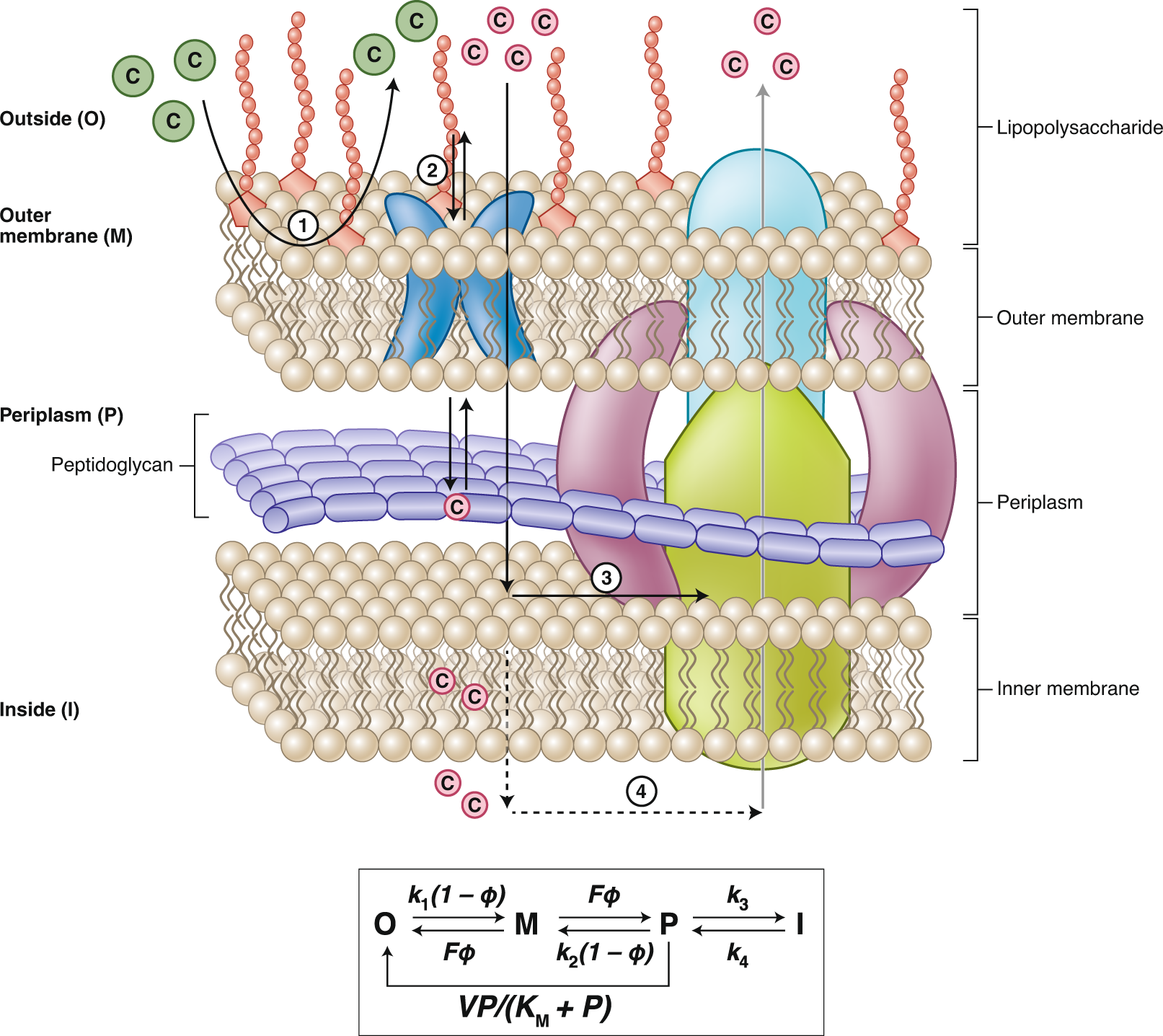
Defining new chemical space for drug penetration into Gram-negative bacteria | Nature Chemical Biology

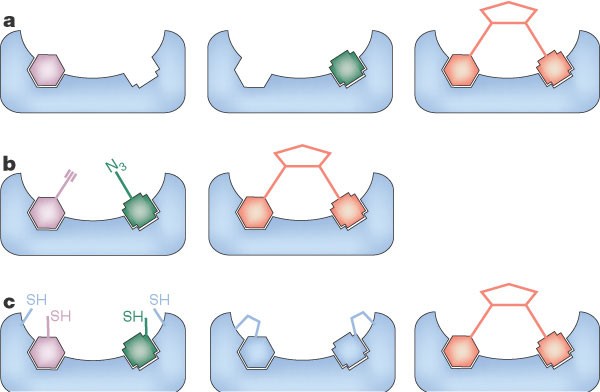
DNA-encoded chemistry: enabling the deeper sampling of chemical space | Nature Reviews Drug Discovery
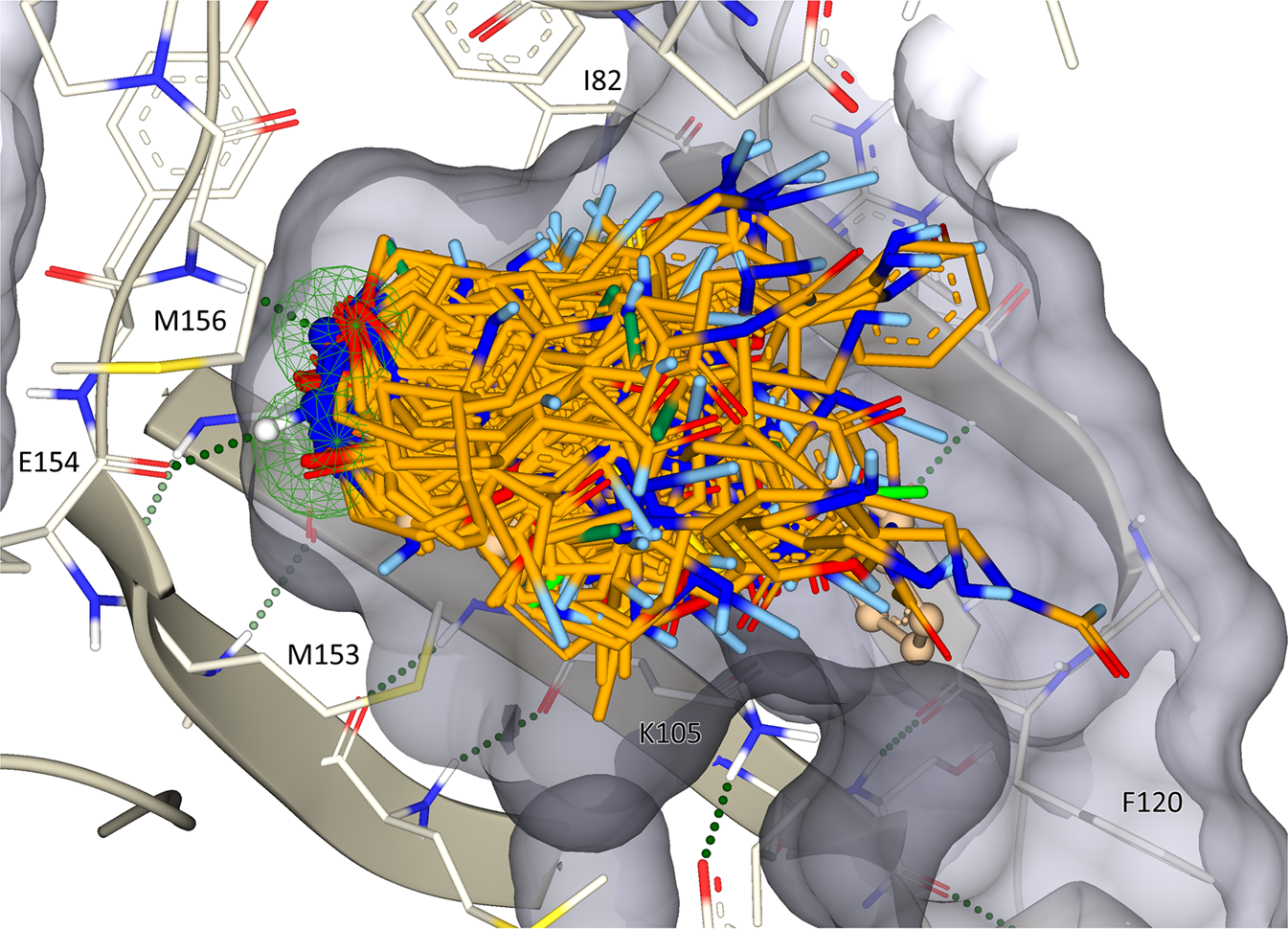
Chemical space docking enables large-scale structure-based virtual screening to discover ROCK1 kinase inhibitors | Nature Communications

Chemical space exploration based on recurrent neural networks: applications in discovering kinase inhibitors | Journal of Cheminformatics | Full Text
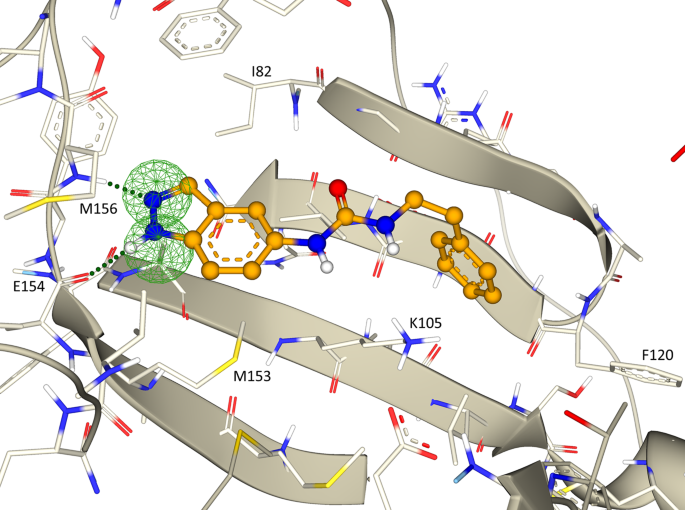
Chemical space docking enables large-scale structure-based virtual screening to discover ROCK1 kinase inhibitors | Nature Communications
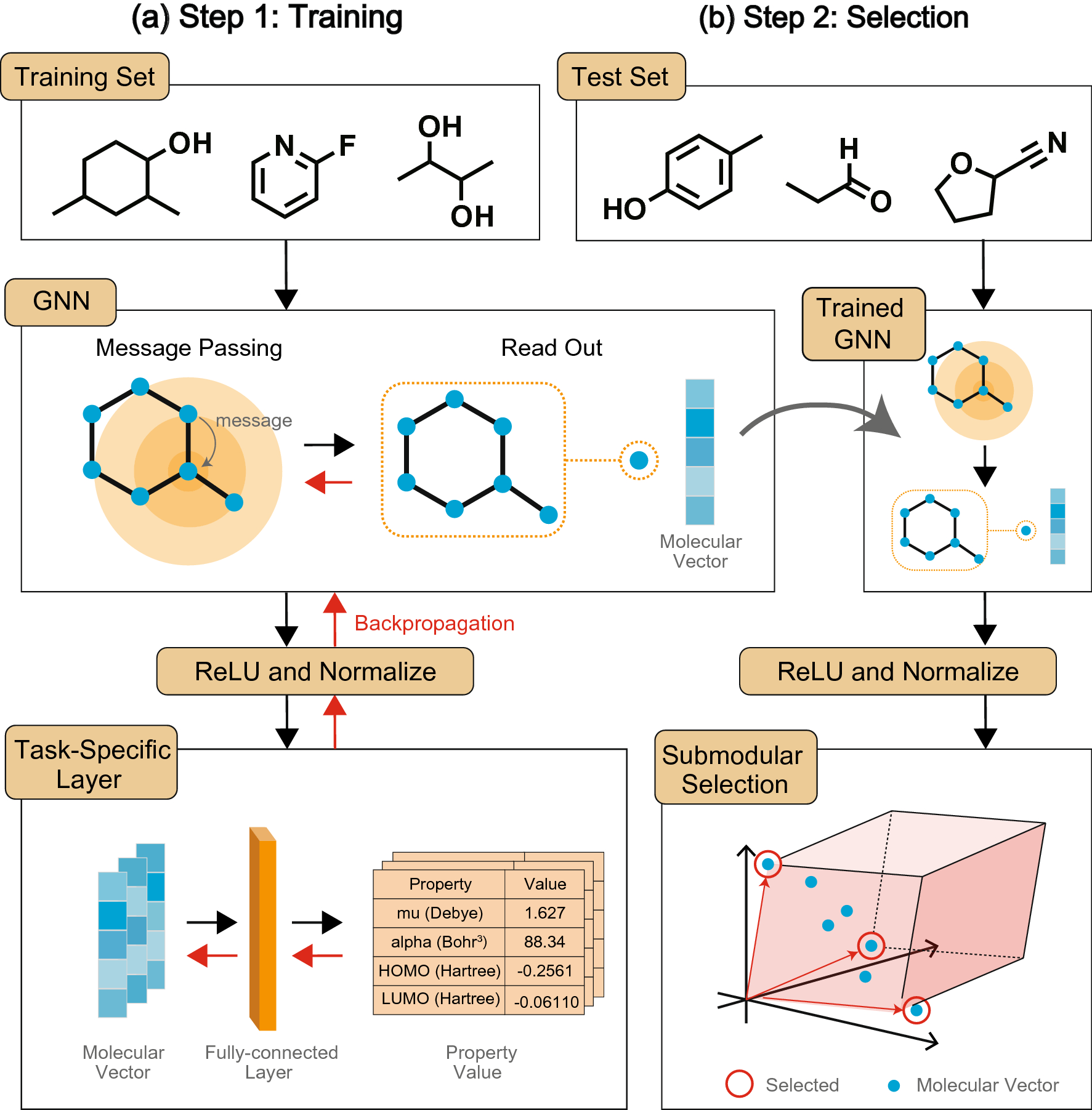
Selecting molecules with diverse structures and properties by maximizing submodular functions of descriptors learned with graph neural networks | Scientific Reports
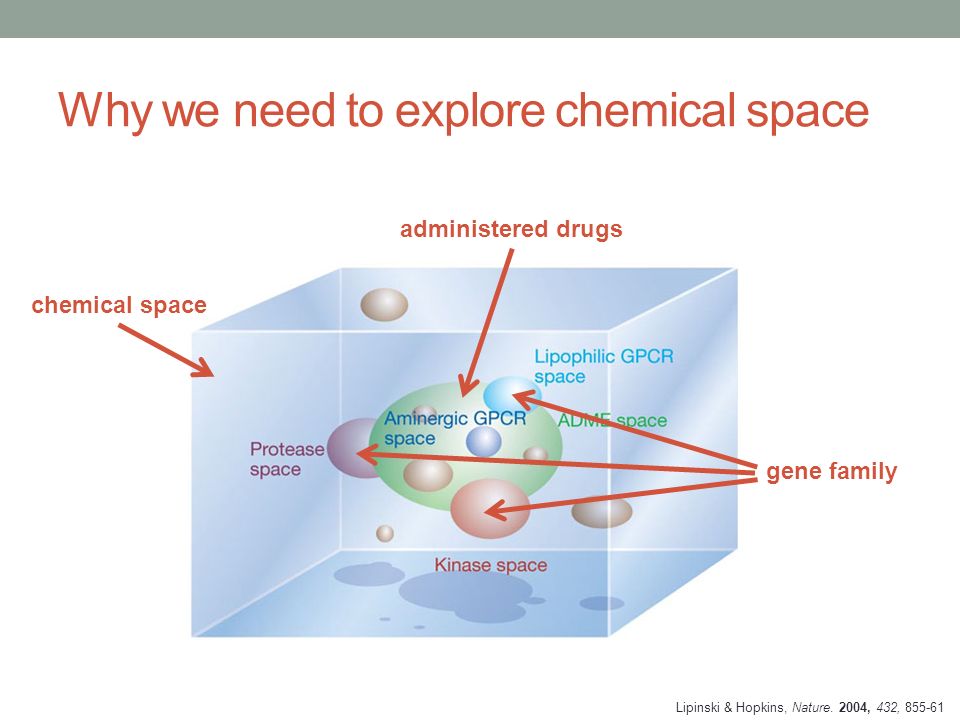
EXPLORING CHEMICAL SPACE FOR DRUG DISCOVERY Daniel Svozil Laboratory of Informatics and Chemistry. - ppt download